Common Run Submission Errors
When you submit read data to ENA, we store and accession your files within Runs. As part of the submission process, read data files must be uploaded to your Webin account’s FTP directory. After you complete the submission, several validation procedures are applied to the file(s). If validation is successful files are archived, otherwise all account contacts are notified of the error(s).
The errors discussed here typically do not require you to repeat the submission in its entirety. It is usually sufficient to upload a corrected version of the file, possibly updating its MD5 value. Once you correct your submission, it can take 24 hours or longer for this to be registered and for the error notifications to cease.
The common error types are described here; if you have been notified of an error, please find it in this list and follow the instructions:
A couple of general-purpose solutions are described too:
If your problem is not described on this page, or you are not clear on the solution, please contact us through our support form.
Error: Invalid File Checksum
If this error occurs, you will receive an email containing something similar to the below:
FILE_NAME | ERROR | MD5 | FILE_SIZE | DATE | RUN_ID/ANALYSIS_ID
mbr_depth_05.bam | Invalid file checksum | 594934819a1571f805ff299807431da4 | 895557023 | 20-DEC-2016 14:02:50 | ERR1766300
The Problem
The checksum is a means of checking a file has been uploaded in its entirety. It is a 32-character string calculated from the file, and is unique to that file. We recalculate the MD5sum after you complete your submission and confirm that it matches the value you registered. Therefore, if the upload procedure fails to deliver the full file, this will be evident from the checksum. You may have calculated this value previously and included it in your submission: you can see the value you registered in the notification email, as is the case above. Alternatively, if you used the graphical Webin File Uploader program, the MD5 will have been calculated automatically for you.
The error could indicate any of the following: 1. A failure in the file transfer process as described 2. The MD5 value was not registered in lower case letters 3. The wrong MD5 value was registered in the first place
The Solution
Depending on the exact cause of the error, there are two possible solutions. Please see Appendix: Correcting An MD5 Value for information on how you can calculate the MD5 value of your local copy of the file and determine whether it matches the originally registered value. If they match, you are looking at a corrupted file error. If they do not match, you are dealing with an incorrectly registered value. In either case, please refer to the relevant section below.
Corrupted File: Upload Again
If you recalculate the MD5 value of you file locally and it matches the value you registered, it is likely the file upload was incomplete or corrupt. You therefore need to reupload the file. Once this is done, the file should be automatically accepted within 24 hours with no further action taken.
Please see Appendix: Re-Uploading Your File for information on how to replace the uploaded file.
Wrong MD5 Value Registered: Register a New One
If you recalculate the MD5 value of your local file and it does not match the value you registered, this will be the root of the problem.
You will need to re-register the MD5 value. Please see Appendix: Correcting An MD5 Value for information on how to do this.
If you have many runs to update, you may wish to do this programmatically, by submitting corrected XML versions of your runs. View our pages on Programmatic Run Updates to learn more about this.
Error: Number Of Lines Is Not A Multiple Of Four
You will receive an email resembling the below if this error occurs:
FILE_NAME | ERROR | MD5 | FILE_SIZE | DATE | RUN_ID/ANALYSIS_ID
SOC9/MCONS1_R1.fq.gz | File content missing or malformed, Number of lines in fastq is not multiple of 4 | c2f8455c1a024cfb96a6c91f5d71f534 | 1358349886 | 01-DEC-2016 03:12:35 | ERR1755094
The Problem
This error is specific to FASTQ files: each read record in such a file should comprise exactly four lines, none of which should be blank. Whitespace characters may only occur in the identifier line and are discouraged. The FASTQ format is as follows:
@<run accession>.<spot index> [<spot name>][/<read index>]
<bases>
+
<phred qualities, ASCII encoded starting with '!' (33)>
If your file does not match this format it may have started incorrectly formatted, or may have become corrupted in the upload process.
The Solution
You can replicate the check we run on your file locally from the command line:
$ zcat <file>.fq.gz | grep '[^[:space:]]' | wc -l
This command will output the amount of lines in the file, after removing any blank lines. Please note that blank lines are considered a violation of the FASTQ format, as empty reads are not informative.
If the line is count divisible by four, it is likely the file was corrupted during upload and should be reuploaded. If it is not divisible by four, you should discover why, correct your file and reupload.
Note
If you reformat your file and then reupload it, you will also need to re-register the checksum. See the Appendix: Correcting An MD5 Value for information on how to do this.
Error: Invalid File Content
If an invalid file content error occurs, you will receive an email with the below message:
FILE_NAME | ERROR | MD5 | FILE_SIZE | DATE | RUN_ID/ANALYSIS_ID
UFMG-CM-Y030_R1.fastq.gz | Invalid file content | 2da9b9c9bb8833c14b103e0de123829c | 137298909 | 13-JUN-2020 12:51:29 | ERR2299965
The Problem
Submitted files content has not been validated. There are two possible reasons for this happening:
There was a delay during the processing of your run files causing the error to be reported
The content of the read file submission does not meet our validation standard
The Solution
First please wait 1 week for the files to be processed. Sometimes your Run records may be affected by a delay in processing your submission, and you will receive the error email before the run processing is completed.
If, after 1 week, the run record is still failing, you can check and update the file content, and then re-upload your run file. Please refer to our guide for Accepted Read Data Formats to help identify your issue.
If you cannot find a problem with your read file contents, please contact our helpdesk.
Updating a run file
Note that if you uploaded the original file to a subdirectory in your submission area, you must also upload the new file to this subdirectory. The processing pipeline expects to see the file for your run in the originally specified location, so this must be maintained. You can check what path the pipeline is expecting to see by referring to the ‘FILE_NAME’ field of the error message: this will contain the full path. See Appendix: Re-Uploading Your File for information on how to correctly upload your file.
Note
If you reformat your file and then reupload it, you will also need to re-register the checksum. See the Appendix: Correcting An MD5 Value for information on how to do this.
Error: Missing File
If a missing file error occurs, you will receive the below message:
FILE_NAME | ERROR | MD5 | FILE_SIZE | DATE | RUN_ID/ANALYSIS_ID
UFMG-CM-Y030_R1.fastq.gz | Missing file | 2da9b9c9bb8833c14b103e0de123829c | 137298909 | 13-JUN-2020 12:51:29 | ERR2299965
The Problem
Submitted files occasionally go missing and must either be replaced or resubmitted.
The Solution
You should reupload the file to your submission area. Note that if you uploaded the original file to a subdirectory in your submission area, you must also upload the new file to this subdirectory. The processing pipeline expects to see the file for your run in the originally specified location, so this must be maintained. You can check what path the pipeline is expecting to see by referring to the ‘FILE_NAME’ field of the error message: this will contain the full path. See Appendix: Re-Uploading Your File for information on how to correctly upload your file.
Appendix: Correcting An MD5 Value
If the MD5 value registered for your read file is incorrect, you can supply a corrected version. To do this:
Log into the Webin Portal
Go to the ‘Run Files Report’
Enter the erroneous run accession into the search field
Identify the run, then select its ‘Action’ box and choose ‘Edit run XML’
You will be presented with the run in XML format: find the
<FILE>elementsThere may be one or more
<FILE>elements, e.g. if you have submitted paired FASTQ files. Find the relevant one by reference to its ‘filename’ valueRemove the ‘checksum’ value for the errored file(s) and enter the correct value. The checksum is highlighted in pink in the image shown below:
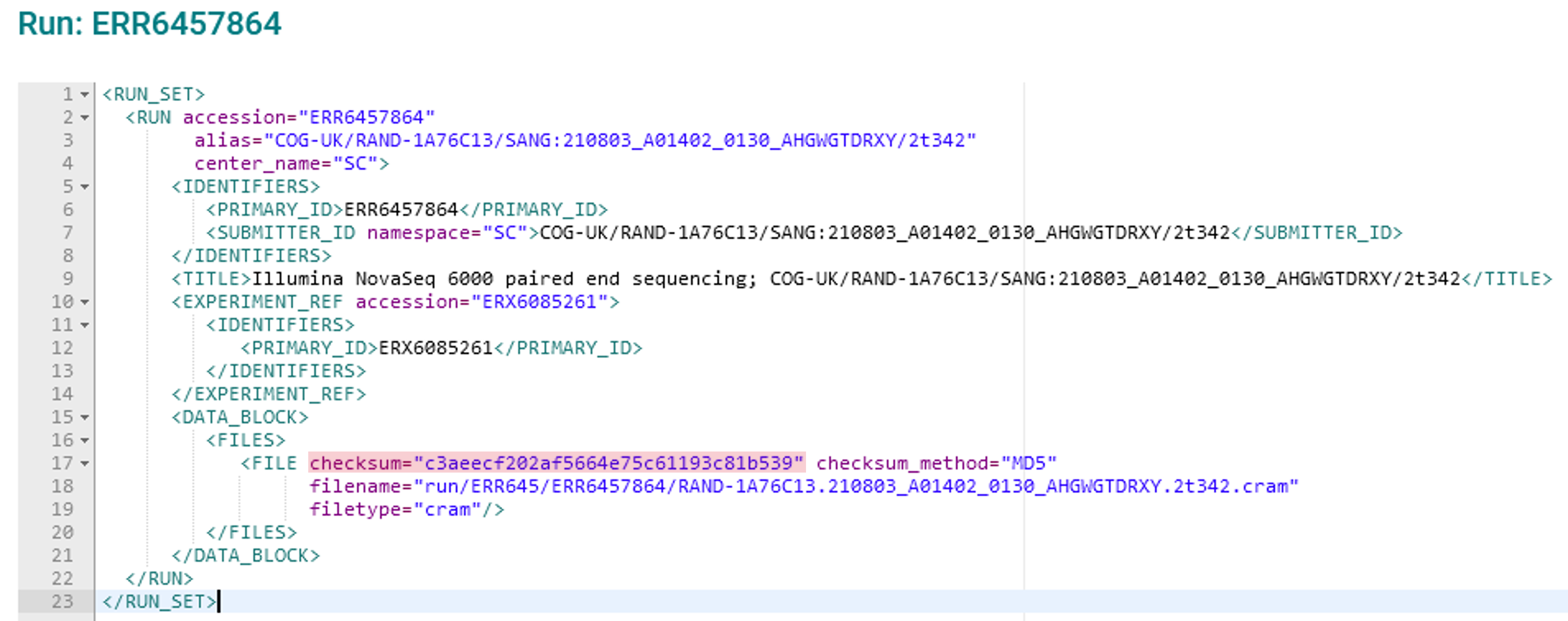
When editing the checksum value, change only the 32-digit string: do not remove the quotation marks, the word ‘checksum’, or any other parts of the XML
Click ‘Save’ at the bottom of the pop-up
Once completed, your file will be re-validated, usually within 24 hours.
Calculating the MD5 value you need can be done natively from the command line in Mac/Linux. One of the following commands will work, if you supply the correct filename:
$ md5sum mbr_depth_05.bam
594934819a1571f805ff299807431da4 mbr_depth_05.bam
$ md5 mbr_depth_05.bam
594934819a1571f805ff299807431da4 mbr_depth_05.bam
For Windows users, 3rd party tools can be found to calculate MD5 values.
Appendix: Re-Uploading Your File
If your error requires a new version of the file be uploaded, you have two options for this. You should first consider whether your file was originally uploaded to a sub-directory. You can tell by referring to the original error message, looking out for the ‘FILE_NAME’ column. The below error describes a file which was uploaded to a subdirectory:
FILE_NAME | ERROR | MD5 | FILE_SIZE | DATE | RUN_ID/ANALYSIS_ID
SOC9/MCONS1_R1.fq.gz | File content missing or malformed, Number of lines in fastq is not multiple of 4 | c2f8455c1a024cfb96a6c91f5d71f534 | 1358349886 | 01-DEC-2016 03:12:35 | ERR1755094
You can tell this was uploaded to a subdirectory because the actual filename ( MCONS1_R1.fq.gz ) is preceded by a directory name and a ‘/’ character ( SOC9/ ). The replacement file must be uploaded to this same subdirectory, as this is where the processing pipeline expects to find it. Having determined this, refer to the relevant section below.
In either case, you may need to update the MD5 value if the originally registered value was correct for the originally uploaded file. If you need to update the MD5 value, please refer to Appendix: Correcting An MD5 Value.
If Your File Is Not In A Subdirectory
Please view our guidance on the Webin File Uploader. This will conveniently allow you to upload your file to the top level of your submission directory.
If Your File Is In A Subdirectory
You will need to upload your file using FTP Client. There are various options for doing this, described at the linked page.
If using a command line solution: Once you are connected to the FTP server, use the ls command to view the content
of the directory and the cd <directory-name> command to move into the required location.
Once you arrive in the desired directory, proceed to upload the files.
Note
If runs fail due to a user error, and not addressed within 180 days, they will automatically get cancelled.