How to Submit Assemblies
Introduction
To submit genome or transcriptome assemblies to ENA you must also provide some metadata to describe your research project. This helps make your data re-useable and searchable.
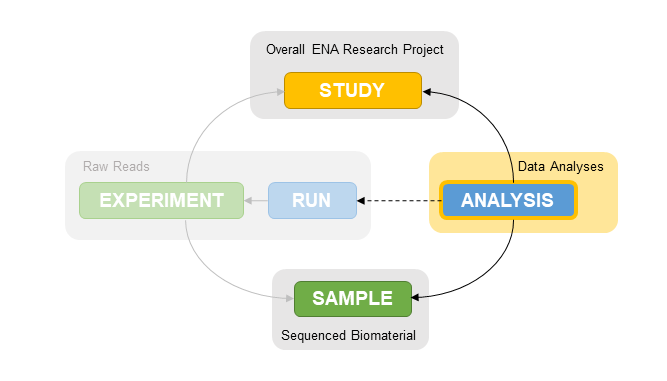
Within ENA, all assemblies are submitted as ‘analysis’ submission objects but are processed differently depending on what type of assembly is submitted.
If you are not yet familiar with the metadata model, please see here for some more information.
As an assembly references ENA sample and study objects, you must submit these before you submit your data. It is also strongly recommended to submit as well as reference any reads associated with the assembly being submitted.
See below for information on how to: register a study within ENA to describe your overall research project, register samples with information on the biological material that was sequenced then assembled, and submit any reads associated with each sample being submitted.
Assembly Levels
Before submitting your assembly, consider the highest level of assembly which has been attained. This will have implications for how you prepare your submission, as well as the accessions you receive at the end.
ENA recognises three assembly levels which describe the highest level of sequence within the assembly. An assembly may contain a mixture of the three sequence types:
Contig: the highest level of assembly is contigs
Scaffold: the highest level of assembly consists of gapped contigs (scaffolds)
Chromosome: the highest level of assembly includes assembled chromosomes
Note that ‘chromosome’ should here be understood as a general term for a range of complete replicons, including chromosomes of eukaryotes, prokaryotes, and viruses, as well as organellar chromosomes and plasmids. All of these may be submitted within the same chromosome-level assembly.
Please also note that contig and scaffold level assemblies can both be updated to higher level assemblies after submission. You cannot update to a lower level assembly, however, and you cannot add functional annotation if none was present in the first submission.
Files For Genome Assembly Submissions
File requirements for a genome assembly submission depends on the assembly level and are specified using a manifest file. The set of files required for genome assembly submissions are listed in the following table:
Assembly Level |
File Requirements |
Additional Information |
|---|---|---|
Contig |
1 Manifest file |
Defines essential metadata |
0-1 FASTA files |
For unannotated assemblies |
|
0-1 EMBL flat files |
For annotated assemblies |
|
Scaffold |
1 Manifest file |
Defines essential metadata |
0-1 FASTA files |
For unannotated assemblies |
|
0-1 EMBL flat files |
For annotated assemblies |
|
0-1 AGP file |
For scaffold instructions from contigs |
|
Chromosome |
1 Manifest file |
Defines essential metadata |
0-1 FASTA files |
For unannotated assemblies |
|
0-1 EMBL flat files |
For annotated assemblies |
|
1 Chromosome list file |
Indicate which sequences represent which ‘chromosomes’ |
|
0-1 Unlocalised list files |
For chromosomes containing unlocalised sequences |
|
0-1 AGP file |
To submit unplaced contigs and indicate which scaffolds/contigs are assembled to form each chromosome |
Accessions
As all assemblies in ENA are submitted as ‘analyses’, for each assembly submission, Webin will report a unique accession number that starts with ERZ. For most assemblies, this accession number is for internal processing only and will not be visible in the browser. As a result, for most assemblies you will receive additional post-processing accession numbers starting with GCA_.
Always make a note of any accessions you receive as these are the unique identifiers for each of your submissions to ENA.
The ERZ accession can be used to access information on the progress of the internal processing of each assembly through the Webin Portal. You can also use this service to see the assigned chromosome, contig, and scaffold accessions. Please follow the Webin Portal link to learn more about this. See individual submission guidelines for information on what accessions you will receive for each assembly type.
In alignment with INSDC partners, SARS-CoV-2 assemblies will not be assigned a GCA_ accession. For these assemblies, sequence accessions will continue to be assigned and the ERZ records will also be available in the browser to provide a point of access for the submitted file(s).
Submission Options
Genome and transcriptome assemblies can only be submitted using the Webin-CLI submission interface. For an overview of how to use this, please see the documentation on Webin-CLI Submission.